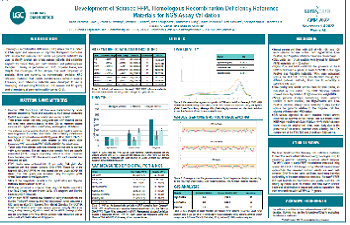
bTMB
Development of Blood TMB reference materials for validation of ccfDNA-based targeted NGS assays that measure tumor mutational burden in patient blood samples
Tumor mutational burden (TMB) is a promising biomarker for predicting positive patient response to immune checkpoint inhibitors. TMB measurements can be determined using genomic DNA extracted from FFPE-preserved tissue biopsy samples.
ctDNA
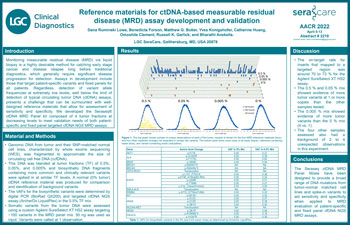
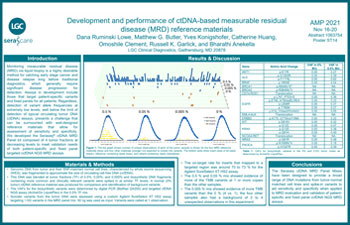
Development and performance of ctDNA based measurable residual disease reference materials
Monitoring measurable residual disease(MRD) via liquid biopsy is a highly desirable method for catching early stage cancer and disease relapse long before traditional diagnostics, which generally require significant disease progression for detection.

cfDNA
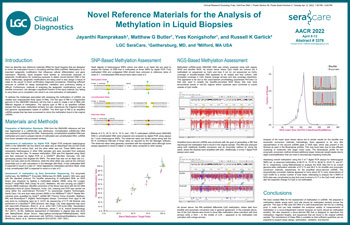
Reference Materials for the Analysis of Methylation in Circulating Cell-Free DNA
Methylation is an important epigenetic modification that influences cellular differentiation and gene expression. Recently, liquid biopsies have started to incorporate analyses of epigenetics modifications for screening purposes to detect cancer-derived DNA in the blood.
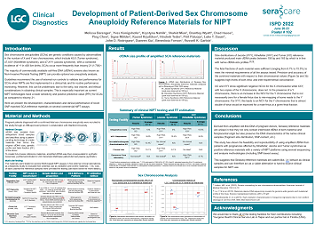
NIPT
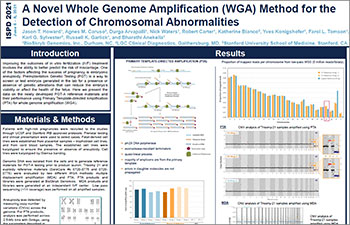
A Novel Whole-genome Amplification (WGA) Method for the Detection of Chromosomal Abnormalities
Improving the outcomes of in vitro fertilization (IVF) treatment means being able to better predict the risk of miscarriage. One of the factors affecting the success of pregnancy is embryonic aneuploidy.

NIPT
Prenatal Clinical Samples enable development of QC reference materials for prenatal and preimplantation genetic testing
Advancing prenatal testing requires an adequate supply of reference materials with clinical and genetic diversity. Using patient remnant samples for assay controls is not a viable or scalable option.
TMB
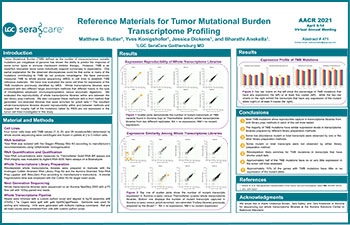
Reference Materials for Tumor Mutational Burden Transcriptome Profiling
Tumor Mutational Burden (TMB) has shown the ability to predict the response of some tumor types to immune checkpoint inhibitor therapy. However, TMB is an imperfect biomarker. We have previously measured TMB by whole exome sequencing (WES) in cell lines to establish TMB reference materials. We have now evaluated the same cell lines for expression of the TMB mutations previously identified by WES.
RNA
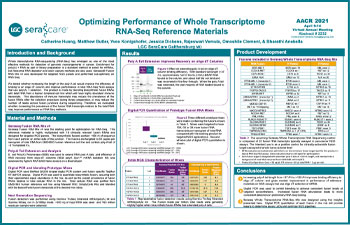
Optimizing Performance of Whole Transcriptome RNA-Seq Reference Materials
Whole transcriptome RNA-sequencing (RNA-Seq) has emerged as one of the most effective methods for detection of genomic rearrangements in cancer. Enrichment for poly(A) + RNA as part of library preparation is a standard method to select for mRNAs, but ribosomal RNA depletion and exon capture methods are also used. Seraseq® Fusion RNA Mix v4 was developed for targeted NGS panels and performed sub-optimally on RNA-Seq.

NIPT
Development of Matched Maternal-Fetal NIPT Reference Materials
How to assure quality of NIPT tests and concordance of test results between various platforms limited availability of clinical samples which can be used for proficiency testing, assay validations, and run controls?

DNA
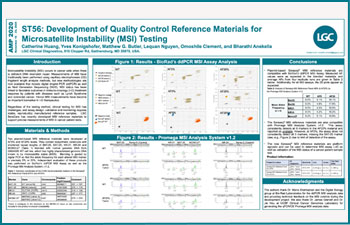
Development of Quality Control Reference Materials for MSI Testing
Microsatellite Instability (MSI) occurs in cancer cells when there is deficient DNA mismatch repair.
TMB
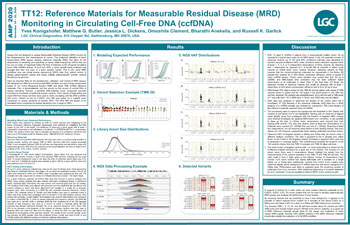
Reference Materials for Measurable Residual Disease (MRD) Monitoring in Circulating Cell-Free DNA
Presented at AMP 2020, this poster describes RMs for the development, validation, and control of MRD assays. We designed our RMs for...
TMB
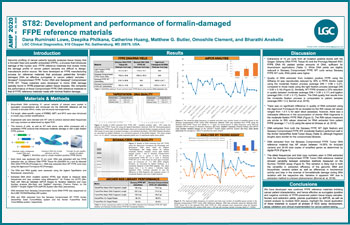
Development and performance of formalin-damaged FFPE reference materials
FFPE reference materials that closely mimic the damage profile of cancer patient samples are difficult to design, manufacture and/or source. We have developed an FFPE manufacturing process for reference materials...
bTMB
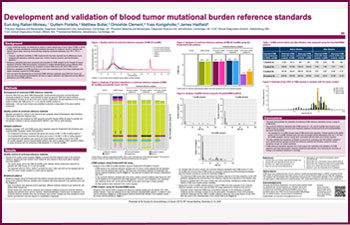
Development and validation of blood tumor mutational burden reference standards
We announced the availability of a range of human diseased cell line based Seraseq® Blood TMB (bTMB) reference controls to support the validation and precise measurements of plasma-derived blood tumor mutational burden...

TMB
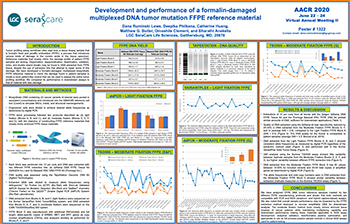
Development and performance of a formalin-damaged multiplexed DNA tumor mutation FFPE reference material
FFPE reference materials that closely mimic the damage profile of cancer patient samples are difficult to design, manufacture and/or source. We have developed an FFPE manufacturing process for reference materials...

TMB
Development of reference material for blood tumor mutational burden measurement
Although most TMB measurements are made from genomic DNA that has been extracted from formalin-fixed paraffin embedded tissue sections, some diagnostic tests attempt to measure TMB in circulating tumor DNA (ctDNA) extracted from blood samples; an approach referred to as blood TMB (bTMB).

TMB
Reference Materials for Measurable Residual Disease (MRD) Monitoring
The analytical validation of liquid biopsy-based assays that attempt to monitor for the disappearance and reemergence of cancer can be challenging due to the need for reference materials...

NIPT
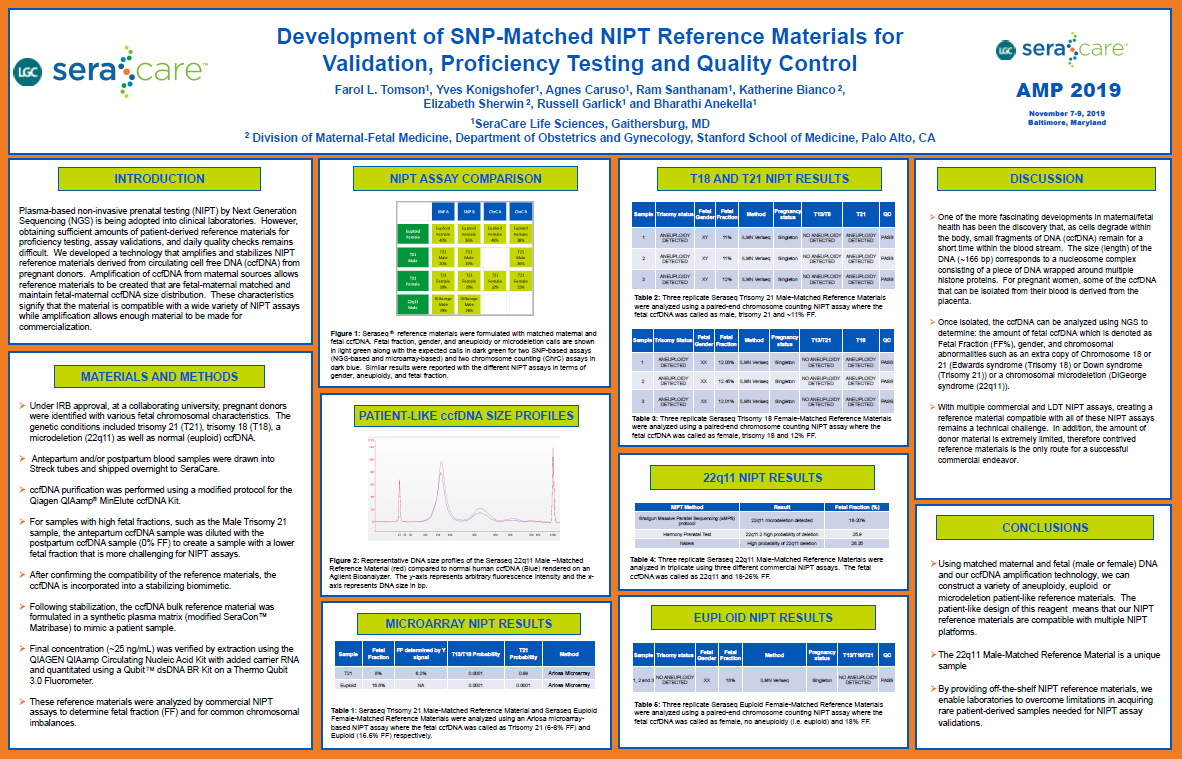
Development of SNP Matched NIPT Reference Materials for Validation, Proficiency Testing and Quality Control
Plasma based non invasive prenatal testing (NIPT) by Next Generation Sequencing (NGS) is being adopted into clinical laboratories. However, obtaining sufficient amounts of patient derived reference materials for...

TMB
Improving and Standardizing TMB Assay Performance
Determining and using the most effective and safest treatment is of great importance in cancer disease management. Recently, a potential biomarker has been identified in immunotherapy: tumor mutational burden (TMB), an assessment of the number of relevant mutations in a tumor.

NTRK
Development of NTRK Reference Materials for Global Assay Standardization
Genomic cancer testing is a powerful tool for identifying the individual and often complex genomic alterations underlying carcinogenesis and cancer progression.
ctDNA
Multi-Center Evaluation of Circulating Tumor DNA Assays
Measurements of circulating tumor DNA (ctDNA) hold great potential to detect residual disease and monitor therapeutic response and tumor evolution. However, there are many technical challenges including trace levels of circulating cell free DNA, short DNA fragment sizes, low variant allelic fractions, and the need to detect all variant types.
RNA Fusion
Consistent Performance of Highly Multiplexed RNA Fusion Reference Materials Across Different NGS Assays in a Multi-Lab Study
Fusion detection is an important part of cancer disease management, and there is an increasing need for highly multiplexed reference materials to cover mutations which may be rare or difficult to obtain.

SRS
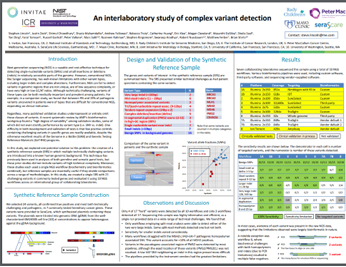
Interlaboratory Assessment of Complex Variant Detection
In this study, an international group of collaborating laboratories explored the creation of a synthetic reference sample (SRS) in which multiple technically challenging variants are introduced into a known human genomic background. This single SRS with 23 variants was evaluated using 10 NGS workflows.
ctDNA
A Comprehensive, Targeted NGS Method that Rapidly and Accurately Detects ctDNA Variants at 0.1% Frequency in Plasma Samples
See how this targeted NGS method is able to distinguish a low-frequency ctDNA signal from background noise in plasma cell-free DNA using a streamlined PCR-based workflow.
ctDNA
New Technology to Generate Commutable and Comprehensive ctDNA Reference Materials for NGS
The need for commutable and comprehensive ctDNA reference materials is evident from the increasing number of liquid biopsy diagnostics and comprehensive panels on the market that are accompanied by reports of discordant results.

NIPT
Highly Stable and Commutable NIPT Reference Materials for Validation, Proficiency Testing and Quality Control
Plasma-based DNA next-generation sequencing (NGS) diagnostics for non-invasive prenatal testing (NIPT) has exploded in popularity in recent years. However...
HCV RNA
Technology to Produce Non-Infectious Recombinant Virus as Reference Materials for Unculturable or Highly Dangerous Viral Pathogens
Diagnostic laboratories and test developers need to design, manufacture, and validate assays for pathogenic viruses and this requires stable, reproducibly manufactured positive reference materials.

NGS
Flexible Tools For The Development And Performance Verification Of Customized Target Enrichment Panels
Implementation of next-generation sequencing for variant identification and discovery presents difficulties both in the selection of genomic loci for inclusion in a panel, as well as the ability
RNA Fusion
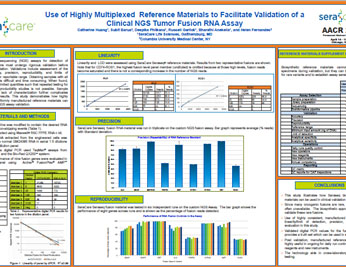
Use of Highly Multiplexed Reference Materials to Facilitate Validation of a Clinical NGS Tumor Fusion RNA Assay
NGS Tumor Fusion RNA Assay Next-generation sequencing (NGS) assays for detection of tumor RNA fusions must undergo rigorous validation before clinical implementation.

Zika
AccuSpan™ Zika Linearity Panel Spans the Dynamic Range of Assays and Allows Evaluation of Analytical Sensitivity
In response to the Zika virus outbreak, several PCR-based assays have been developed and approved under the Emergency Use Authorization (EUA).

NGS
Multiplexed Next-Generation Sequencing Reference Materials for Testing of Inherited Disorders
Next-generation sequencing (NGS) has rapidly advanced the genetic testing for inherited disorders.

ctDNA
A Commutable Circulating Tumor DNA (ctDNA) Reference Material
Commutability is an important aspect of reference materials and relates to how well the materials can mimic the natural analyte.
NGS
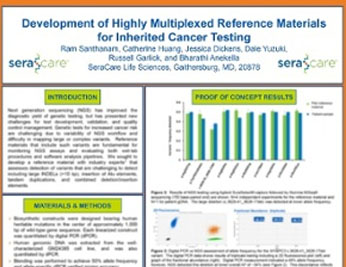
Development of Highly Multiplexed Reference Materials for Inherited Cancer Testing
Next Generation sequencing (NGS) has improved the diagonistic yield of genetic testing, but has presented new challenges for test development, validation and quality control management.