Reference Standards for Tumor Profiling Analysis

LGC SeraCare is the industry leader in NGS-based Reference Standards
LGC SeraCare’s portfolio of Seraseq® oncology reference products are industry standards for use in a broad range of NGS-based tumor profiling and immuno-oncology (I-O) assay workflows (see, e.g., Figure-1), used in disease diagnosis and I-O treatment outcome decisions. We offer RNA and DNA-based reference standards, in all formats (purified gDNA, RNA and FFPE) for use as positive sample controls in all phases of NGS assay needs – development, validation and clinical applications.
![]()
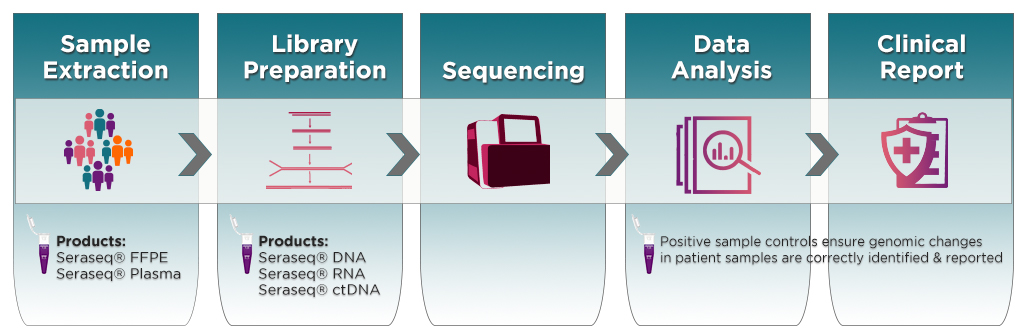
Figure-1: A sample-to-report workflow for targeted NGS assays and applications of Seraseq controls for ensuring assay quality and accuracy of test results.
DNA Standards
Developing and optimizing somatic mutation assays is a difficult task, with variations in the amount of tumor cellularity and potential sources of variability across the NGS workflow. Successful assays require accuracy throughout the entire process, from sample DNA purification and quantitation, to library construction and template preparation, through bioinformatics parameters and variant annotation.
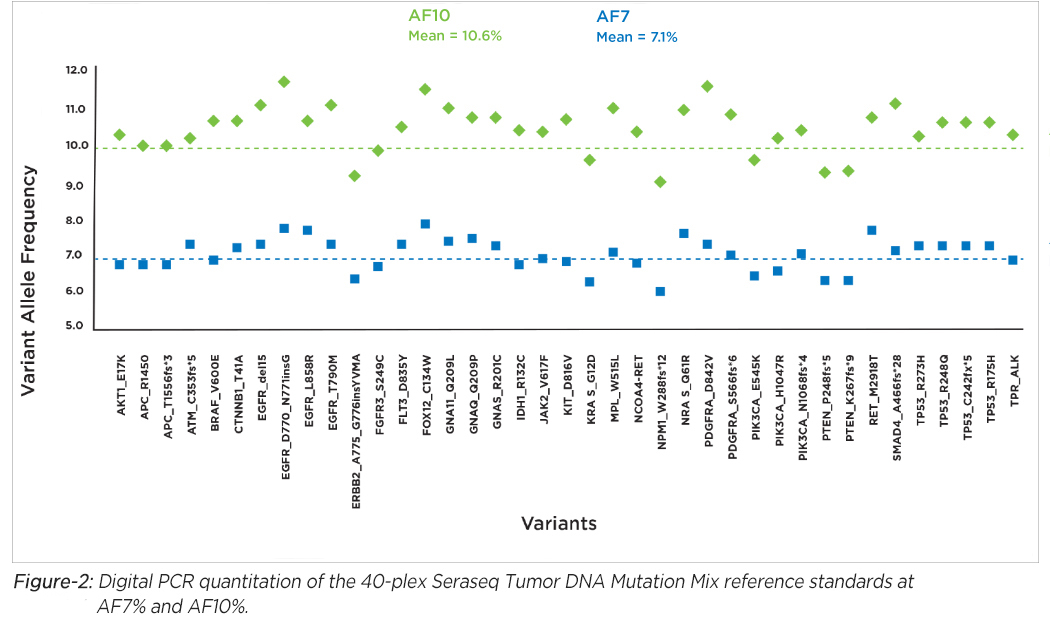
Seraseq's DNA products offer mutation analysis of 40 clinically-relevant variants (SNVs, INDEL, SVs) blended at AF7% and AF10% in a single reference sample (see Figure-2). We also offer DNA reference standards for analyzing CNV amplifications of clinical relevance in some cancer diseases.
List of DNA genes
| AKT1 | CTNNB1 | FLT3 | GNAS | KRAS | NRAS/CSDE1 | RET |
| APC | EGFR | FOXL2 | IDH1 | MPL | PDGFRA | SMAD4 |
| ATM | ERBB2 | GNA11 | JAK2 | NCOA4-RET | PIK3CA | TP53 |
| BRAF | FGFR3 | GNAQ | KIT | NPM1 | PTEN | TPR-ALK |
Contact us to learn more about our DNA Products
RNA Standards
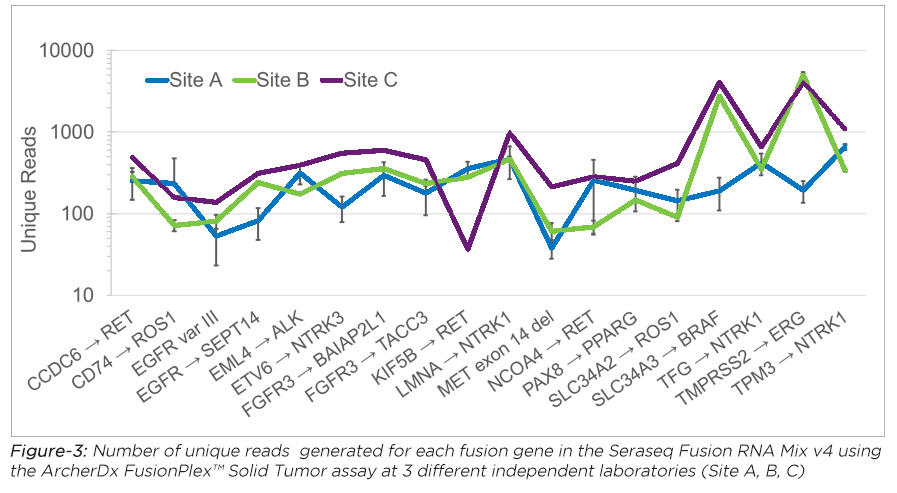
LGC SeraCare has developed the most comprehensive panel of highly multiplexed Seraseq Fusion RNA reference materials to match these new highly muliplexed targeted RNA-based NGS assays. These products are offered in both a purified RNA format for ease of use in evaluating library preparation and sequencing, as well as a full-process FFPE reference material to assess performance starting with pre-analytical steps.
List of RNA fusion genes
| CCDC6-RET | FGFR3-BAIAP2L1 | PAX8-PPARG1 | IRF2BP2-NTRK1 | PAN3-NTRK2 |
| CD74-ROS1 | FGFR3-TACC3 | SLC34A2-ROS1 | SQSTM1-NTRK1 | BTBD1-NTRK3 |
| EGFR Variant III | KIF5B-RET | SLC45A3-BRAF | AFAP1-NTRK2 | |
| EGFR-SEPT14 | LMNA-NTRK1 | TFG-NTRK1 | NACC2-NTRK2 | |
| EML4-ALK | MET ex14 Skipping | TMPRSS2-ERG | QKI-NTRK2 | |
| ETV6-NTRK3 | NCOA4-RET | TPM3-NTRK1* | TRIM24-NTRK2 |
For more on RNA standards read our blog posts on: Multi-Lab FFPE Fusion and Multi-Lab Fusion RNA Mix assessments for NGS assays.
| Seraseq Solid Tumor RNA Products |
| Seraseq® FFPE Tumor Fusion RNA v4 Reference Material |
| Seraseq® Fusion RNA Mix v4 |
| Seraseq® FFPE NTRK Fusion RNA Reference Material |
| Seraseq® NTRK Fusion RNA Mix |
Whole Transcriptome RNA Sequencing Mix
The new Seraseq Whole Transcriptome RNA Sequencing control is composed of fusion RNA transcripts typically assayed in pan-cancer NGS assays. The intended use is as a positive control for clinically-actionable fusion targets assayed at whole transcriptome level, and includes the following characteristics:
- A panel of 22 RNA fusion targets at >10,000 copies/µL.
- Each fusion target is designed with extended poly-A tails of >100 nt length, and blended into a background of total RNA extracted from GM24385 cell line (WT).
- Product Configuration: Conc = 50 ng/µl; Fill volume = 20 µl; total RNA ~ 1 µg per vial.
Fusion Gene List
|
# |
RNA Fusion |
5' Partner |
3' Partner |
|
1 |
BCR-ABL1 | BCR exon 14 | ABL exon 2 |
|
2 |
CCDC6-RET | CCDC6 ex 1 | RET ex 12 |
|
3 |
CD74-ROS1 | CD74 ex 6 | ROS1 ex 34 |
|
4 |
EML4-ALK | EML4 ex 13 | ALK ex 20 |
|
5 |
ETV6-ABL1 (transcript 1) | ETV6 exon 4 | ABL exon 2 |
|
6 |
ETV6-ABL1 (transcript 2) | ETV6 exon 5 | ABL exon 2 |
|
7 |
ETV6-NTRK3 | ETV6 ex 5 | NTRK3 ex 15 |
|
8 |
FGFR3-TACC3 | FGFR3 ex 17 | TACC3 ex 11 |
|
9 |
KIF5B-RET | KIF5B ex 24 | RET ex 11 |
|
10 |
LMNA-NTRK1 | LMNA exon 11 | NTRK1 exon 11 |
|
11 |
LMNA-NTRK1 | LMNA ex 2 | NTRK1 ex 10 |
|
12 |
MEF2D-CSF1R | MEF2D ex 7 | CSF1R ex 11 |
|
13 |
MET ex 14 Skipping | MET ex 13 | MET ex 15 |
|
14 |
NACC2-NTRK2 | NACC2 exon 4 | NTRK2 exon 13 |
|
15 |
NCOA4-RET | NCOA4 ex 8 | RET ex 12 |
|
16 |
PML-RARα | PML exon 6 | RARα intron 2 |
|
17 |
RUNX1-RUNX1T1 | RUNX1 exon 6 | RUNX1T1 exon 2 |
|
18 |
SLC34A-ROS1 | SLC34A2 ex 4 | ROS1 ex 34 |
|
19 |
SLC45A3-BRAF | SLC45A3 ex 1 | BRAF ex 8 |
|
20 |
TCF3-PBX1 | TCF3 exon 16 | PBX1 exon 3 |
| 21 | TMPRSS2-ERG | TMPRSS2 ex 1 (5' UTR) | ERG ex 2 |
| 22 | TPM3-NTRK1 | TPM3 ex 7 | NTRK1 ex 9 |
Contact us to learn more about our RNA Products
![]()
Immuno-oncology Standards
TMB Products
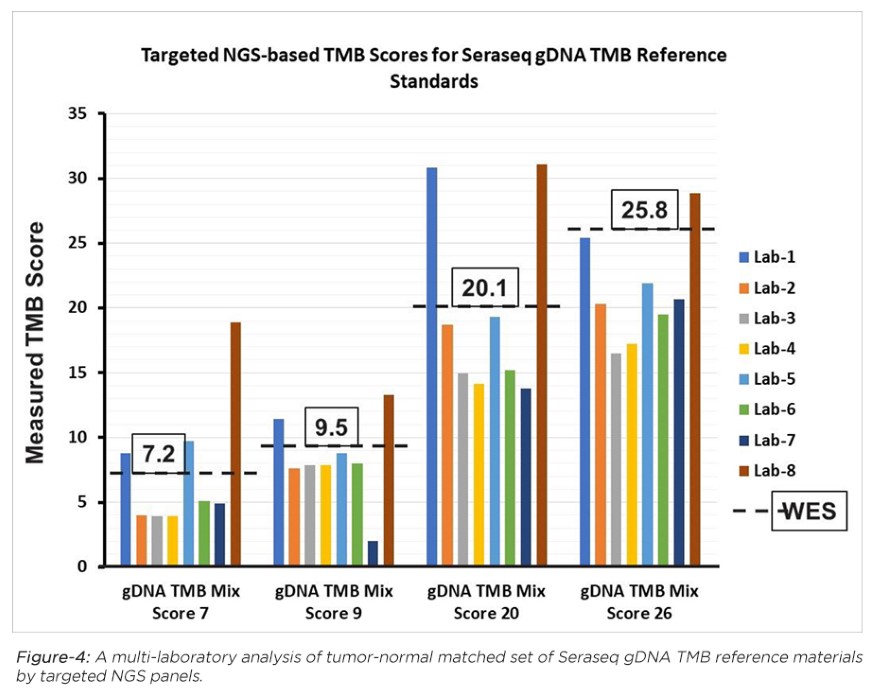
LGC SeraCare’s industry-first TMB reference standards are derived from tumor-normal matched human cell lines, and characterized by whole exome sequencing (WES) and a TMB analysis pipeline to determine their empirical TMB scores (see Table-1). Developed in collaboration with industry experts and analyzed by a range of NGS targeted panels and analysis pipelines (see, e.g., Figure 4), these reference standards provide ground-truth information for accurate determination of cancer patient TMB scores.
Table-1: Seraseq TMB reference standards and WES-derived TMB scores
|
Products |
TMB Scores* (gDNA) |
TMB Scores* (FFPE) |
| Seraseq® TMB Score 7 | 7.2 ± 0.2 | 7.15 ± 0.4 |
| Seraseq® TMB Score 9 | 9.5 ± 0.4 | 7.52 ± 1.3 |
| Seraseq® TMB Score 13 NEW! | 12.6 ± 0.02 | 12.1 ± 0.3 |
| Seraseq® TMB Score 20 | 20.1 ± 0.2 | 18.59 ± 0.5 |
| Seraseq® TMB Score 26 | 25.8 ± 0.5 | 22.8 ± 3.6 |
*Based on Whole Exome Sequencing (WES)
Microsatellite Instability (MSI) Standards
LGC SeraCare has developed microsatellite instability (MSI) reference materials containing the key mono- and dinucleotide biomarkers typically analyzed in molecular MSI assays – NR-21, NR-24, BAT-25, BAT-26 and MONO-27. These markers are blended at two different allele frequency levels – AF5% and AF20% - to support MSI assay LoD determinations and accurate MSI detection. These products are offered as tumor-normal matched pairs, with variants precisely quantitated by digital PCR as well as qPCR/CE against a highly characterized genomic DNA from a background WT cell line (GM24385) determined as microsatellite stable (MSS).
| Seraseq MSI Products |
|
| Seraseq MSI Reference Panel Mix AF5% | Seraseq MSI Reference Panel Mix AF20% |
| Seraseq gDNA MSI-High Mix NEW! | |
![]()
Scientific Resources
To support our commitment and value we have selected assets we hope you find valuable. CLICK HERE to download some of the scientific content for our Seraseq® tumor profiling and immuno-oncology reference standards. After completing the form, you will receive an email with a link to all downloads.