NGS Resources

NIPT
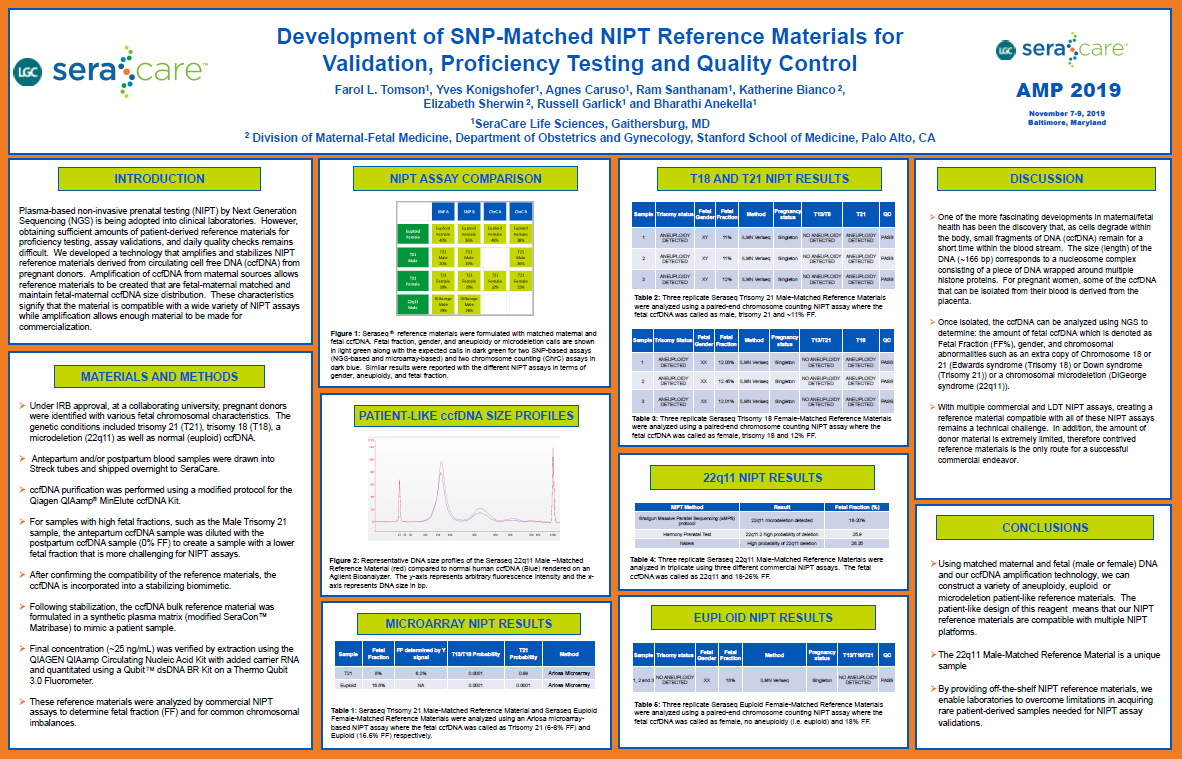
Development of SNP Matched NIPT Reference Materials for Validation, Proficiency Testing and Quality Control
Plasma based non invasive prenatal testing (NIPT) by Next Generation Sequencing (NGS) is being adopted into clinical laboratories. However, obtaining sufficient amounts of patient derived reference materials for...

TMB
Improving and Standardizing TMB Assay Performance
Determining and using the most effective and safest treatment is of great importance in cancer disease management. Recently, a potential biomarker has been identified in immunotherapy: tumor mutational burden (TMB), an assessment of the number of relevant mutations in a tumor.

TMB
Optimizing TMB Use in Cancer Research and Care: The Friends of Cancer Research TMB Harmonization Effort
Harmonization of methods to quantify TMB will facilitate robust biomarker development and optimize clinical utilization and treatment decision-making. Friends aims to better understand the impact of assay variation on clinical outcomes, align standards, and define best practices for TMB assessment.

TMB
Target the Tumor Mutational Burden and Pursue Harmonization
Friends aims to better understand the impact of assay variation on clinical outcomes, align standards, and define best practices for TMB assessment. Listen as Dr. Mark Stewart discusses the role of the tumor mutational burden (TMB) in cancer research and the need for harmonization throughout the process.

NTRK
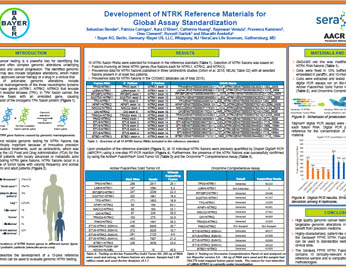
Development of NTRK Reference Materials for Global Assay Standardization
Genomic cancer testing is a powerful tool for identifying the individual and often complex genomic alterations underlying carcinogenesis and cancer progression. SeraCare has developed reference material that contains 15 clinically-relevant NTRK fusions in a single reference sample.
ctDNA
New Technology to Generate Commutable and Comprehensive ctDNA Reference Materials for NGS
The need for commutable and comprehensive ctDNA reference materials is evident from the increasing number of liquid biopsy diagnostics and comprehensive panels on the market that are accompanied by reports of discordant results.

Validation eBook
Next-Generation Sequencing Assay Validation Guide
A practical guide to validating a clinical NGS assay with real-world examples from leading clinical genomics experts.
ctDNA
A Comprehensive, Targeted NGS Method that Rapidly and Accurately Detects ctDNA Variants at 0.1% Frequency in Plasma Samples
See how this targeted NGS method is able to distinguish a low-frequency ctDNA signal from background noise in plasma cell-free DNA using a streamlined PCR-based workflow.
Validation
Validate Your Clinical Genomics Assay Without Crushing Your Budget or Sanity
In this video, Drs. Greg Tsongalis and Russell Garlick will review how to employ modern NGS QC tools to accelerate the development of your clinical genomics assay for less than it costs with traditional materials and methods.

NGS Implementation
Best Practices for Validation, Monitoring, and Controls
A panel of clinical genomics experts provides an overview of the regulatory landscape for laboratory-developed tests. Panelists discuss how clinical genomics labs can and should ensure the analytical and clinical validity of their tests amid an uncertain regulatory climate.

Liquid Biopsy
Expert Perspectives on Liquid Biopsy Clinical Implementations
Drs. Tony Godfrey, Peggy Gulley, and Greg Tsongalis provide expert perspectives on what it takes to achieve full clinical potential for ctDNA assays.
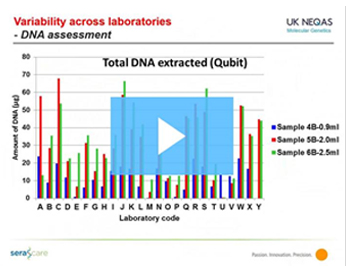
Case Study
Enabling Clinical Genomics with Highly Multiplexed and Patient-Like Reference Materials
Dr. Sandi Deans, Director of UK NEQAS for Molecular Genetics, presents a case study of how a global EQA organization ensures the accuracy and consistency of a clinical genomics application.

Assay Development
How to Develop a Clinical NGS Assay Without Losing Your Mind or Your Shirt
This guide contains best practices to help you develop an NGS-based clinical genomics assay and bring it online as quickly and cost-effectively as possible. Overcome the top three assay development challenges faced by clinical laboratories.
